Here you can find ongoing work at REBLAB. Our most important finding so far is the connection between a type 1 pullulanase and Lactobacillus crispatus glycogen metabolism. Find the post here, and accompanying protocols and data here. Find an introductory lecture to the field and my work here. I also send out a newsletter, if you want stay up to date subscribe here
Research proposal: genetic factors of host glycogen metabolism and its relationship with the vaginal microbiome.
After visiting the Keystone conference I was wondering if genetic factors of the hostess could be related to her vaginal microbiome. Janneke van de Wijgert pointed me towards the HELIUS study, that has both SNP data and vaginal microbiome available for a few hundred participants. I wrote a proposal to compare these datasets. I hope we will be able to perform this study this year. If you are working on something comparable, let me know and perhaps we can join forces! If you have any other proposals of promising SNP’s that we should take into account in the analysis, I would be very interested to hear about this.
Summary Evidence is accumulating that glycogen released by endometrium and the cervicovaginal epithelium functions as an important carbohydrate for vaginal bacteria. Luminal glycogen is found to vary both with hormonal status and with the bacterial make-up of the reproductive tract. Here we propose to study whether host genetic factors that are involved in glycogen metabolism regulation show correlation with vaginal microbial signatures.
The multi-ethnic HELIUS cohort of Amsterdam is uniquely positioned for this analysis [1]. The vaginal microbiome of several participants was characterized previously [2], showing a variation in Lactobacillus-dominated microbiota and Lactobacillus-depleted dysbiotic states. This latter microbial state is associated with an increased risk of preterm labor and acquisition of sexually transmitted infection. Of a subset of these women (346 in total) genome wide SNP data will become available this year. We propose to analyze whether women with Lactobacillus-dominated vaginal microbiota are significantly more likely to carry certain alleles in the most prominent SNP’s of the TCF7L2 gene that are associated with type 2 diabetes. This preliminary analysis data could inform novel studies within this same HELIUS cohort to study the correlations between host vaginal glycogen synthesis, lactate concentration and vaginal microbiome.
Hypothesis: Glycogen functions as a carbon source for Lactobacillus species colonizing and acidifying the human vagina. We hypothesize that genetic changes in the glycogen synthesis pathway and its regulatory factors can affect the ability of Lactobacillus species to acidify the vagina and prevent changing to a bacterial vaginosis microbial state. Moreover, unpublished experiments have shown that bacterial members of this dysbiotic state such as Gardnerella and Prevotella also benefit from glycogen to grow to high numbers, produce biogenic amines (odor), cause inflammation (recruiting target cells for HIV to the mucosal surfaces), desialylation of the mucosa and exfoliation of the vaginal epithelium (discharge). Lower glycogen stores may partially explain why certain women with this microbial state have less symptoms, such as odor, discharge and inflammation, than other women.
Background During reproductive years glycogen is synthesized by the tissues of the reproductive tract [3, 4] and is essential for early embryo implantation and development. Glycogen shed into the vaginal lumen functions as a carbon source for vaginal lactobacilli such as Lactobacillus crispatus [5, 6] and iners but also for Gardnerella vaginalis and Prevotella bivia (unpublished data). Glycogen levels are reduced in women who have Lactobacillus-depleted microbial states [7] which could be explained by the interplay between host factors (glycogen synthesis) and bacterial factors (glycogen breakdown) [8].
It has been well-established that ethnicity is an important factor determining the odds of a woman to have bacterial vaginosis. Women of African descent are found to have more Lactobacillus iners than Lactobacillus crispatus and are more often colonized by a Lactobacillus-depleted community [9-11]. These women often have symptoms such as discharge and odor, and are at higher risk of acquiring sexually transmitted infection [12, 13], are more likely to have persistent HPV colonization [14, 15] and are more likely to give birth prematurely [15-17].
There has been considerable effort to understand the variation of health outcomes and symptoms amongst women with comparable vaginal microbiota. Most studies were directed at understanding this variation from a bacterial perspective, for instance by looking at genetic variation amongst bacterial isolates of Gardnerella vaginalis [18, 19] and Lactobacillus crispatus [6, 20]. We believe that the dataset we propose to analyze provides a unique opportunity to also take host factors into consideration.
Variations in glycogen metabolism due to host genetic factors may affect both hepatic glycogen accumulation as well as endometrial and vaginal glycogen accumulation. For a long time it was thought that progesterone was the main controlling hormone for glycogen synthesis in the endometrium [21, 22]. A recent study has now found that the progesterone/glycogen link is only indirect. Insulin is the hormone that inactivates glycogen synthase kinase 3β thereby activating glycogen synthase, similarly in liver tissue as well as in endometrial cells [23].
These commonalities lead to our hypothesis that there may be a genetic relationship between bacterial vaginosis and type 2 diabetes. In genome-wide association studies into genetic risk factors for type 2 diabetes several SNP’s have been identified. The most robust and consistent SNP’s are found in the TCF7L2 gene. Certain SNP’s in this transcription factor (elsewhere referred to as a TCF/LEF or TCF7) confer up to twofold increased risk of developing type 2 diabetes [24]. TCF7L2 knockout mice had reduced glycogen stores in the liver [25]. People carrying the risk allele synthesized less insulin in response to a glucose challenge test [26, 27]. This lower insulin production may have consequences for glycogen accumulation in the reproductive tract. Less glycogen synthesized means less carbohydrate for vaginal Lactobacillus species to acidify and Gardnerella to cause symptoms, odor and discharge. As of yet the ethnic differences in type 2 diabetes (where African ethnicity are at higher risk) were not able to be traced back to any SNP’s. The link between one specific TCF7L2 SNP’s (rs7903146) and type 2 diabetes was found in various regions of the world.
This would be the first study into host genetic factors and vaginal colonization.. The strength that this analysis offers is that it does not only include people from European heritage but includes a variety of groups with Asian, African and European roots. Clues from this study may inform bigger studies where we could look at the influence of genetic factors on lactate and glycogen taking into account the different ethnic groups and vaginal colonization patterns. If glycogen metabolic commonalities are found between diabetes and BV, this opens up a wide range of new treatment options for BV and related conditions, and possibly novel applications for established diabetes drugs including amylase inhibitors, insulin and metformin.
Proposal
To study the SNP’s of the TCF7L2 gene and their relationship to vaginal microbiome. The HELIUS cohort will this year have both datasets available for 346 women. The vaginal microbiome of these women was previopusly studied and the authors are willing to share these data too. We propose to look at three SNP’s of the TCF7L2 gene, specifically rs7903146, rs 4506565 and rs12255372 and study whether their relationship with having a Lactobacillus-deplete (case) or Lactobacillus-dominated (control) microbiome. In case the study cohorts permit (and the odds ratio), we could differentiate further into ethnicity. In case we find a strong signal that would warrant further study, we could look at glycogen concentration, lactate concentration, vaginal microbiome and ethnicity. There are many more vaginal swabs available and SNP data for 12,000 participants in total.
Collaboration and transparency This is an Open Kitchen Science project, meaning that the aim is to maximize collaboration and transparency. All data files, methods and results, either positive or negative, will be released and published prior to peer review (either on a preprint-server or on blog). Regular information about proceedings will be shared (including this proposal, without financial details) on blogs.
1. Snijder, M.B., et al., Cohort profile: the Healthy Life in an Urban Setting (HELIUS) study in Amsterdam, The Netherlands. BMJ Open, 2017. 7(12): p. e017873.
2. Borgdorff, H., et al., The association between ethnicity and vaginal microbiota composition in Amsterdam, the Netherlands. PLOS ONE, 2017. 12(7): p. e0181135.
3. Milwidsky, A., Z. Palti, and A. Gutman, Glycogen metabolism of the human endometrium. J Clin Endocrinol Metab, 1980. 51(4): p. 765-70.
4. Mirmonsef, P., et al., Glycogen Levels in Undiluted Genital Fluid and Their Relationship to Vaginal pH, Estrogen, and Progesterone. PLoS One, 2016. 11(4): p. e0153553.
5. Hertzberger, R.Y., A. Brandt, and R. Kort, Carbohydrate active enzymes in Lactobacillus crispatus – a possible link between the pullulanase gene and growth on glycogen. Figshare, 2018.
6. van der Veer, C., et al., Comparative genomics of human Lactobacillus crispatus isolates reveals genes for glycosylation and glycogen degradation: Implications for in vivo dominance of the vaginal microbiota. bioRxiv, 2018: p. 441972.
7. Mirmonsef, P., et al., Free glycogen in vaginal fluids is associated with Lactobacillus colonization and low vaginal pH. PloS one, 2014. 9(7): p. e102467.
8. Vaneechoutte, M., The human vaginal microbial community. Res Microbiol, 2017. 168(9-10): p. 811-825.
9. Fettweis, J.M., et al., Differences in vaginal microbiome in African American women versus women of European ancestry. Microbiology, 2014. 160(Pt 10): p. 2272-2282.
10. Ma, B., L.J. Forney, and J. Ravel, Vaginal microbiome: rethinking health and disease. Annual Review of Microbiology, 2012. 66: p. 371-389.
11. Ravel, J., et al., Vaginal microbiome of reproductive-age women. Proceedings of the National Academy of Sciences of the United States of America, 2011. 108 Suppl 1: p. 4680-4687.
12. Gosmann, C., et al., Lactobacillus-Deficient Cervicovaginal Bacterial Communities Are Associated with Increased HIV Acquisition in Young South African Women. Immunity, 2017. 46(1): p. 29-37.
13. Sewankambo, N., et al., HIV-1 infection associated with abnormal vaginal flora morphology and bacterial vaginosis. Lancet (London, England), 1997. 350(9077): p. 546-550.
14. Kero, K., et al., Association of asymptomatic bacterial vaginosis with persistence of female genital human papillomavirus infection. Eur J Clin Microbiol Infect Dis, 2017. 36(11): p. 2215-2219.
15. Brown, R.G., et al., Establishment of vaginal microbiota composition in early pregnancy and its association with subsequent preterm prelabour rupture of the fetal membranes. Translational Research, 2018.
16. Donati, L., et al., Vaginal microbial flora and outcome of pregnancy. Archives of Gynecology and Obstetrics, 2010. 281(4): p. 589-600.
17. Martius, J. and D.A. Eschenbach, The role of bacterial vaginosis as a cause of amniotic fluid infection, chorioamnionitis and prematurity–a review. Archives of Gynecology and Obstetrics, 1990. 247(1): p. 1-13.
18. Schellenberg, J.J., M.H. Patterson, and J.E. Hill, Gardnerella vaginalis diversity and ecology in relation to vaginal symptoms. Res Microbiol, 2017. 168(9-10): p. 837-844.
19. Janulaitiene, M., et al., Phenotypic characterization of Gardnerella vaginalis subgroups suggests differences in their virulence potential. PLoS One, 2018. 13(7): p. e0200625.
20. France, M.T., H. Mendes-Soares, and L.J. Forney, Genomic Comparisons of Lactobacillus crispatus and Lactobacillus iners Reveal Potential Ecological Drivers of Community Composition in the Vagina. Appl Environ Microbiol, 2016. 82(24): p. 7063-7073.
21. Jaffe, R.C., D.M. Stevens, and H.G. Verhage, The effects of estrogen and progesterone on glycogen and the enzymes involved in its metabolism in the cat uterus. Steroids, 1985. 45(5): p. 453-62.
22. Mimori, H., et al., Effect of progestogen on glycogen metabolism in the endometrium of infertile patients during the menstrual cycle. Fertil Steril, 1981. 35(3): p. 289-95.
23. Flannery, C.A., et al., Insulin Regulates Glycogen Synthesis in Human Endometrial Glands Through Increased GYS2. J Clin Endocrinol Metab, 2018. 103(8): p. 2843-2850.
24. Florez, J.C., et al., TCF7L2 polymorphisms and progression to diabetes in the Diabetes Prevention Program. N Engl J Med, 2006. 355(3): p. 241-50.
25. Boj, S.F., et al., Diabetes risk gene and Wnt effector Tcf7l2/TCF4 controls hepatic response to perinatal and adult metabolic demand. Cell, 2012. 151(7): p. 1595-607.
26. Jainandunsing, S., et al., Transcription factor 7-like 2 gene links increased in vivo insulin synthesis to type 2 diabetes. EBioMedicine, 2018. 30: p. 295-302.
27. Loos, R.J., et al., TCF7L2 polymorphisms modulate proinsulin levels and beta-cell function in a British Europid population. Diabetes, 2007. 56(7): p. 1943-7.
Power to “the crispies” – what I learned at the Keystone symposium on the genital microbiome.
In December I visited the long awaited Keystone symposium on the Role of the Genital Microbiome in Reproductive and Sexual Health. It was such an exciting and interesting conference and I learned a lot. Even in the airplane I spent the very last minutes of the flight talking about vaginal bacteria.
The strength of this symposium was the diversity. There were doctors and lab people, students and professors, there were Africans, Europeans, Americans and Asians. But most importantly, they came from different fields: HIV, preterm birth, etc. These people normally would not be at the same conference, they live parallel scientific lives publishing in their own journals and attending their own meetings. But the bacteria of the vagina bring them together. It has become crystal clear that the vaginal flora (or “microbiota”) have a very strong say in the various questions of these fields, such as who is prone to HIV infection (sexual health) and who is at high risk to deliver their baby too soon (reproductive health).
These people sat down in a big conference room together for four days, with in total 22 hours of presentations and about ten hours of more random wine-infused scientific chatter around 70 square feet of results printed on big poster boards.
The collective results were striking to say the least. One study that turned up in several presentations a group of 236 young South-African women were followed for about a year. Just as a reminder: in certain areas of Africa women start out with 0% HIV at 14 years old and in the following six years about 60% gets infected with HIV. In certain communities being seropositive is the norm. In this particular study, on the FRESH cohort in a township in Kwazulu Natal, a group of women were followed for about two years, 13% of all women got infected with HIV during this time period. But the risks of getting HIV-infected were not the same for all of them. The subgroup that had a vaginal flora mostly made up of a bacterium called Lactobacillus crispatus exactly 0 girls were infected. These bacteria seemed to almost behave like condoms (although please please still use condoms).
Find the paper here: https://linkinghub.elsevier.com/retrieve/pii/S1074-7613(16)30519-2
This same picture returned in literally every presentation that followed. Women with Lactobacillus species always have better health outcomes, and in most studies women with Lactobacillus crispatus do a bit better than Lactobacillus iners. The only exception I can think off is Candida infection that occurs regardless of the vaginal microbiome or in some studies a bit more often in women with vaginal lactobacilli.
Jeanne Marazzo, the opening speaker of the symposium, showed a picture of the vaginal lactobacilli a protective superpower. A dangerous watchdog that keeps out the bad guys. It was fascinating for me, yet again, to see this one Lactobacillus species beyond any doubt, in virtually every talk at the conference, in any cohort, be the good guy. I confirms that it is important and worthwhile to focus on these bacterial species because they mean so much for women’s health.
What happens if you don’t have these crispies (as I heard them refer to during the conference) or you lose them in the course of your life? Well, one of the most prevalent vaginal bacteria is a species called Lactobacillus iners (either pronounced like “ainers” pronounced like “I nurse” or “inners” like in “inner circle”). Clearly, this is the less preferred Lactobacillus. A bacterium with two faces: women that have this bacterium as their most abundant species seem less well protected and more likely to get infected or when pregnant deliver their babies too early. However, Lactobacillus iners is still preferred over having no Lactobacillus at all. You could see iners as just not a very good watchdog. Maybe he doesn’t bite? Or maybe he is secretly friendly with the bad guys? Or maybe he is just careless: leaving windows and doors open to let the bad guys in? All questions up for debate.
Douglas Kwon, who was the senior author on the previously mentioned study on HIV, showed data that sketch a rough storyline of vaginal microbiome. Most women start out with “crispies”, if they change to a different bacteria, it’s most often iners, and if they change from iners their more likely to switch to a flora that has very few lactobacilli but lots of other unwanted species such as Gardnerella, Prevotella and Megaspaera. These bacteria are not regarded as real bad guys (like Salmonella or Chlamydia) that will directly make you sick, although some believe they are. Clearly, you still really need to have unprotected sex with an infected partner before you get infected with Chlamydia or HIV.
Having no lactobacilli but these bacteria like Gardnerella is regarded as a “state” and referred to as “dysbiosis”. Although in many African groups it is more common than having Lactobacillus. Most women wouldn’t even notice that they don’t have good watchdogs around but some have symptoms like discharge and odor. For some it is so serious that it just outright destroys their sex lives, their relationships and their confidence. Even without preterm birth and HIV, this would already be a pretty good reason to study it and try to help these women out.
The immune response
One of the things that I learned more about during the conference is the role of the immune system. Without your lactobacilli, the immune system is more likely to be alarmed. It starts to get ready to attack and reacts as if its fighting off an infectious disease (we call this inflammation: redness, swelling, pain caused by your immune system responding to an infection). This is bad news if you have sex with a partner infected with HIV because HIV viruses infect exactly those cells (T-cells) that are involved in the immune response. So basically, you are calling the victims to the party.
When we talk about the immune system we distinguish between “pro-inflammatory” and “anti-inflammatory” signals: bacteria or chemicals that are the equivalent of someone yelling “fire fire!” to the immune system (pro-inflammatory), or sing a lullaby to the immune system (anti-inflammatory). Both may be going in in the vagina. Species like Gardnerella and Prevotella may already on their own may send alarming signals. And chemicals produced by Lactobacillus may be the equivalent of the lullaby: a comforting signal to the human body that “the good guys” are around.
When your pregnant the story may be a bit different. It could be that these bacteria alone may cause your water to break way too early and give your baby a troublesome start (or worse). One of the mechanisms through which this happens may be inflammation. Several stories in the symposium outline how Lactobacillus species will prevent an inflammation reaction whereas species like Gardnerella will provoke inflammation. Some of the pathways through which our bodies start such an inflammatory response are the same that the pregnant body uses to start labor.
Before I move on to my own research let’s just sit back and look at the weirdness of the vagina. How incredible it is that women really only just have two important watchdogs. In some groups you will find about 80% of women having one of these. This is found nowhere else in the human body and as Jeanne already pointed out: we really need to get rid of “diversity” as a characteristic of a healthy flora (or “microbiome”). In the vagina you want less species not more.
It was really great to meet so many of the heroes of the field. Everyone took a lot of time to answer all questions and everyone was very approachable.
Unfortunately, I cannot share many of the very interesting new aspects of the vaginal microbiome that were presented during the meeting, because a lot of the results are unpublished.
If you want to see the poster I presented, please find it here: https://figshare.com/articles/Lactobacillus_crispatus_growth_on_glycogen_is_dependent_on_its_type_1_pullulanase_gene_variant/7478330
I also gave a 10 minute oral presentation and I used my slides later for a lecture at ARTIS/Micropia. Find the presentation here: https://figshare.com/articles/Introduction_to_vaginal_microbiome_-_lecture/7583312
What’s next?
This year I am planning to make much more time for research. I want to further look at glycogen metabolism of vaginal bacteria and luckily I have met several people at the conference who are working on the same topic and are interested to work together.
The meeting really stresssed the importance of trespassing the boundaries of different fields. I really want to pull the human factor into the lab. Study the presence of glycogen levels and glycogen breakdown enzymes in vaginal swabs. Also, I will continue studying the type 1 pullulanase of L crispatus and regulation of this gene/enzyme. Having seen the talks in Cape Town I think that these basic mechanisms of vaginal metabolism need to be elucidated so we can move on. Deborah Jekel, a very motivated and talented bachelor student is starting this week to help me do just that.
Stay tuned!
News!
Great news! We sent an abstract to the Keystone conference “Role of the Genital Tract Microbiome in Sexual and Reproductive Health” and it was selected for a short talk. So I’ll be presenting the research on stage in Cape Town in December!
This is the abstract:
Lactobacillus crispatus growth on glycogen is dependent on its type 1 pullulanase gene variant.
Rosanne Y. Hertzberger1,2, Alicia Brandt 2,3, Charlotte vd Veer4, Jorne Swanenburg1,5, Remco Kort1,5
1 VU University Amsterdam, Netherlands, 2 REBLAB, Netherlands, 3 University of Groningen, Netherlands, 4 Public Health Service, Amsterdam, Netherlands, 5 TNO, Zeist, Netherlands
Glycogen is an abundant carbohydrate in the vagina of reproductive-age women. Levels of free glycogen vary depending on the makeup of the vaginal microbiota: women with bacterial vaginosis, a common dysbiosis with higher pH and few lactobacilli, generally have lower glycogen levels. This finding has led to the general assumption that lactobacilli depend on this glycogen and use it as a carbon source for colonization and acidification. However, direct evidence for this mechanism has been lacking sofar.
We studied growth, enzymatic activity and metabolite production of a group of 19 Lactobacillus crispatus strains that were isolated from reproductive age women. 13 out of these 19 isolates were able to grow on glycogen whereas 6 did not use glycogen for growth. Starch was used as a proxy to study glycogen breakdown and starch metabolism activity was found in both supernatant as well as in the washed pellets after growth on glycogen. Several strains did not show activity after growth on glucose indicating differential regulation of this metabolic pathway.
A survey of the genomes to track down any carbohydrate active enzymes showed the presence of a putative cell surface associated type 1 pullulanase. The gene is predicted to encode a 140 kDa enzyme consisting of an N-terminal signal peptide, two carbohydrate binding modules, a carbohydrate active module belonging to the GH13_13 family and a C-terminal S-layer associated protein (SLAP) domain. The gene shows high similarity to the extracellular cell wall attached pullulanase previously found in a human gut isolate of Lactobacillus acidophilus.
Although a copy of the pullulanase gene was found in the genomes of all 19 L. crispatus strains, the six non-glycogen consuming strains all carried one of three different mutations in the N-terminal signal peptide sequence, expected to disrupt transcription of at least part of the gene.
Our results show for the first time that certain vaginal isolates of Lactobacillus crispatus are capable of metabolizing one of the most abundant vaginal carbohydrates. We identify a pullulanase that may be essential for this activity. These findings bring us further to understanding the basic mechanisms of Lactobacillus colonization and acidification of the human vagina.
This project is part of REBLAB, an Open Kitchen Science initiative. For more information see www.reblab.org.
Also: a paper with the glycogen results (reproduced in the TNO lab) is under review at Microbiome journal and available in the meanwhile at bioRxiv, find it here https://www.biorxiv.org/content/early/2018/10/13/441972
Observations…
Hi everybody,
This summer I did some preliminary test with 4 strains (2 glycogen consumers, RL_010 and RL_011, and 2 non-consumers RL_009 and RL_026) to see whether maltose is a suitable substrate to be able to compare activity between glycogen consuming and glycogen non-consuming strains of L. crispatus. I also used this experiment to get some additional replicates for Figure 3 of the bigger June blog post (this one). Will be updating this soon. First need to repurchase a GraphPad PRISM serial code 🙁
Preliminary observations of the experiment:
-something went wrong with the dilution of the culture for strain 9 (5 times instead of 10 times diluted), so I won’t be able to use this data point.
-the 2 glycogen consuming strains show starch degrading activity in both pellets as in the supernatants, like I saw before. (could this be the S-layer effect?)
-strains grow well on maltose but the activity is still somewhat repressed (or not induced?) just like on glucose. Next step is to try this on maltotriose to see if this may be a substrate that all strains will grow on and pullulanase activity will also be present.
-suprisingly, one strain (RL_011) does seem to induce (or fails to repress) the pullulanase activity on glucose and on maltose. Is there another level of variation in this aspect?
Experiment wishlist
Back in the lab! Hope to spend a lot of time here in the coming weeks. My program is getting busier in October so I hope I can get a lot done before that. On my experimental wishlist:
-Study Gardnerella vaginalis and Lactobacillus iners growth on glycogen and starch breakdown. For this I need to be able to create more strict anaerobic conditions then the growth protocols I am using now since these species seem to be more sensitive to oxygen. Let’s see if we can arrange that in the lab.
-Further compare glycogen consumers and “non-consuming” strains of L. crispatus. To be able to make this comparison I will need to find growth conditions in which I get expression of the gene, but growth of both consumers and non-consumers. Alicia told me that maltose, maltotriose or maltopentaose may do the trick. The precise mechanisms of this induction of expression (?, or whatever it may be) are interesting and may have in vivo relevance, but have low priority. I just want to compare the strains, but no growth (of the non-consuming strains on glycogen) means no pellet and no bacteria to use as a comparison! I will start with maltose, and have ordered maltotriose. These experiments will also be useful to get more replicates for the growth data in the previous blog.
-Alicia also told me more about the differences between amylose and amylopectin in starch. This may explain why we only saw partial breakdown of starch in activity assay of L. crispatus. If we are able to differentiate between amylose and amylopectin utilization, we may also learn more about the exact activity and ways to use this as a target for therapeutic approaches.
-At some point I really need to make an overview on what is actually known about glycans in the human vagina. What techniques and stains were used in the different studies sofar to characterize the glycans?
Stay tuned!
Carbohydrate active enzymes in Lactobacillus crispatus – a possible link between the pullulanase gene and growth on glycogen
Hello everyone, after moving to Rotterdam and having a baby (welcome Frank!) I am back here to show you the progress I made! The findings I present here are work in progress, but it is time to give a first update. I believe the results I present here are promising on three different levels: 1) content-wise, I think they are a start to understanding glycogen metabolism of Lactobacillus crispatus better 2) I am showing it all: data, methods, results, questions and flaws. Exactly how I envision Open Kitchen Science, (although with a bit of delay, so less “realtime” as I would like) 3) this is the work of a collaboration with another “postacademic” scientist, who wants to contibute to science besides her regular job. Meet Alicia Brandt!
Before diving into the science, a few points of order:
Collaboration: As mentioned before, I am very interested in collaborating and receiving feedback on my blog posts. Please contact me, or leave a remark. I am also interested whether any other labs are working on the same question or have been working on this in the past. If you are (or were ) doing experiments with Lactobacillus crispatus we could find out whether these results are true for more strains (especially using the same medium). Let’s talk! Twitter or Facebook or email rosanne dot hertzberger at gmail dot com
Manuscript: Part of the results presented here will be part of a paper that is currently in preparation by Charlotte van der Veer and Remco Kort and co-authors (including me). I will update this blog post as soon as this paper is published. When the paper is published, the full genomes of the strains will also be released online.
Internship: starting after the summer break I hope to be supervising a student working on this project, together with Jurgen Haanstra at the VU University Amsterdam. Find the description here (lab-site) or here (FigShare). BSc or MSc, please contact me if you are interested!
Warning: The results presented in this blog post are unreviewed and have not been replicated by an independent laboratory.
Open data All data and protocols used can be found in this FigShare collection.
How to cite: Hertzberger, Rosanne; Brandt, Alicia (2018): REBLAB: Carbohydrate active enzymes in Lactobacillus crispatus – a possible link between the pullulanase gene and growth on glycogen. figshare. Collection.
https://doi.org/10.6084/m9.figshare.c.4133819.v3
Short overview of where we are.
I am interested in glycogen metabolism of vaginal microbes. Glycogen is an abundant vaginal source of carbohydrates and varies depending on the vaginal bacterial signature. Roughly spoken: when the vaginal microbiota are dominated by Lactobacillus glycogen is generally higher. One of the overarching questions in the field is whether vaginal lactobacilli can metabolize this glycogen to grow and produce lactic acid. I think this is a pretty big deal. I have had a Google Scholar Alert with the keywords “vaginal glycogen” since a few years and not a week goes by without a paper getting published talking about how this vaginal glycogen is supposedly turned into lactic acid by lactobacilli. In my view, there is no evidence to back up this statement.
During my postdoc at Washington University St. Louis it became clear that the BV-associated Gardnerella vaginalis and Prevotella bivia are capable of growing on glycogen, as well as Lactobacillus iners (often encountered in vaginal microbiota with and without BV). Previously on this blog I showed experiments that Lactobacillus crispatus can grow and acidify using glycogen as a source. Here we zoom in further on the variation in glycogen metabolic capacity of different Lactobacillus crispatus strains, using a group of 23 different Lactobacillus crispatus isolates.
Summary of the findings
A dbCAN survey of the genomes of these strains shows an overview of several carbohydrate-active enzymes possibly involved in glycogen breakdown. One of the genes present in all strains is a putative S-layer linked type 1 pullulanase (GH13_13). Zooming in on this gene it appears that there is strong variation in the N-terminal sequence encoding a putative signal peptide. Only the strains that do not have a mutation in the sequence of this signal peptide show growth on glycogen. 6 out 23 strains that have a mutation in this peptide cannot grow on glycogen. Is this signal necessary for Lactobacillus crispatus to break down the big molecules of glycogen in the vaginal environment? Nothing final yet, but a promising lead to look further into.
Alicia Brandt: dbCan analysis and glycogen-active enzymes
Since I started with REBLAB I encountered several people who shared their own stories about their departure from science and how this affected them. Sometimes these people find the time and energy to still contribute to science parallel to their regular jobs. We start to call ourselves “post-academic” scientists. One of these people is dr. Alicia Brandt (previously Alicia Lammerts van Bueren as she is known in the glycobiology world) who left science last year for a job in a supporting role at the Young Academy at Groningen University. She expressed the strong wish to keep being involved and to share her knowledge and skill. It’s a happy coincidence that her expertise is exactly what I am looking for: glycobiology and even glycogen metabolism of bacteria. I went to meet her in Groningen and had a great time!
Alicia and me on the steps of the Groningen academy building
Lactobacillus crispatus isolates in this blog
Last year I was very lucky to join Prof. dr. Remco Kort at the VU University Amsterdam who had just isolated and sequenced about 30 Lactobacillus crispatus strains from women with and without BV ecology. These women were patients at the GGD facility in Amsterdam and the strains were isolated as part of the thesis work of Joke Dols. MSc student Jorne Swanenburg was responsible for the genomic analysis sofar and Charlotte van de Veer is currently finalizing her thesis and is writing up a paper on carbohydrate metabolism of these strains. Recently, a paper was published where the strains were used as a reference for a possible new vaginal prebiotic. The paper is open access and the isolation is described in materials&methods. At the moment we are analyzing these strains, some appear harder to culture than others. The list of isolates that we are able to maintain in the lab is a work in progress and will be updated later. We were unable to revive RL_005 from its glycerol stock and the sequencing file of RL_022 has some problems.
Alicia offered to use her expertise to help answer some questions surrounding Lactobacillus crispatus glycogen metabolism. First thing she did was to perform an analysis using the so-called dbCAN server, detecting the presence of several glycoproteins in the genomes of a set of isolated Lactobacillus crispatus strains that can potentially be involved in glycogen metabolism.
Find the raw data here (FigShare).
Find the protocol here (FigShare).
Most important findings:
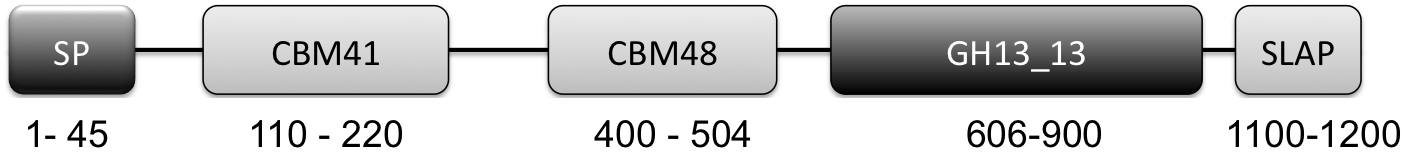
- All L. crispatus genomes contained a putative cell-surface associated (as indicated by SLAP domain) pullulanase Type 1 enzyme (http://www.uniprot.org/uniprot/A0A135Z466) implicated in glycogen degradation. Key features of this enzyme are the presence of a CBM41 and CBM48 and a GH13_13 catalytic domain (Fig 1). More on this gene/protein later in this blog post.
- There is a cluster (operon?) of alpha-glucan degrading enzymes in all crispatus genomes which include a GH65 (phosphorylase?), GH13_20(CBM34) and GH13_31-2 enzyme. Further analysis required to see if they are co-transcribed. (Table 1, genes located in the cluster are indicated with an (a)).
- Other alpha-glucan metabolizing enzymes found within the L. crispatus genomes include: GH13_18, GH13_29, GH13_31, GH31 (see Table 1).
- Only three strains of L. crispatus contained a GH13_18 (RL02, RL09, RL10) (Note: RL_022 contained this enzyme as well, but sequencing file of strain has errors).
- RL_006 is the only strain that does not contain a GH13_29 enzyme.
Table 1: Overview of Alpha-glucan enzymes found in L. crispatus genomes (see www.cazy.org for more info on predicting enzyme activities based on amino acid sequence similarities with known enzymes within a given family)
| Enzyme Family | Proposed Activity | Genomes |
| GH13_13 (CBM41, CBM48) | Pullulanase type I | All except RL31, RL32 |
| GH13_18 | Sucrose phosphorylase | Only RL02, RL09, RL10. RL22* |
| GH13_20 (CBM34)(a) | Pullulanase type III, cyclodextrinase | All |
| GH13_29 | Trehalose-6-phosphate hydrolase | All except RL06 |
| GH13_31-1 | oligo-alpha-1,6-glucosidase | All |
| GH13_31-2 (a) | oligo-alpha-1,6-glucosidase | All |
| GH31-1 | alpha-glucosidase | All |
| GH31-2 | alpha-glucosidase | All |
| GH65 (a) | Maltose phosphorylase | All |
*RL22 genome needs to be resequenced (problems with sequencing file)
(a) Constitutes part of a cluster of enzymes, possibly an operon.
N-terminal signal peptide of the type 1 pullulanase gene (GH13_13) corresponds with growth on glycogen.
Glycogen is a prevalent potential carbon source in the vagina of reproductive age women. Lactobacillus crispatus and Lactobacillus iners-are the most frequently encountered species vaginally. They are generally assumed to be responsible for the low pH and high lactate concentrations- but it is unclear what sugar source they use for lactic acid production.
(To be fair: there are many assumptions and uncertainties here. To name a few: there is only circumstantial evidence that vaginal lactic acid is of bacterial origin. The same counts for the human origin of vaginal glycogen. Actually, the fact that we are dealing with glycogen and not with a different glycan is not all that well established since most studies used a PAS stain and an alpha-glucosidase. It could well be that we are dealing with a different glycan. However, I am cutting a few corners here and will assume that lactate is of bacterial origin and the glycan is in fact glycogen, produced by the hostess herself.)
Previously it was reported that at least a subset of vaginal lactobacilli (jensenii, gasseri and johnsonii) were unable to breakdown glycogen, but this study did not look at glycogen metabolism of crispatus and iners. I presented on this blog some evidence that the DSM strain of L crispatus is capable of growth and acidification (producing lactic acid) on glycogen as the carbon source.
This possible L. crispatus glycogen metabolism could be an important player in the acidification of the human vagina, and the health benefits that are associated with a vaginal community dominated by L. crispatus. Needless to say, we would like to know more about it.
Why the type 1 pullulanase had my interest – the role of serendipity
This gene had had my attention already since I started working with a set of 4 Lactobacillus crispatus strains during my postdoc in St Louis at the Lewis lab, WUSTL School of Medicine. These strains were MV-1A-US, MV-3A-US, JV-V01 and 125-2-CHN. I found that two of these (MV-1A-US and MV-3A-US) were able to grow on glycogen as a carbon source whereas two others (JV-V01 and 125-2-CHN) were not. To find a possible genetic origin of these differences I did a blastp analysis using several enzymes as a query that were known to be involved in glycogen (or starch) breakdown. Those were: glgX of E. coli, sap of Streptococcus agalactiae (see paper and sequence ), SusB of Bacteroides theta and the amylase (amyE) of Bacillus subtilis. Three out of the four genes showed no full-length copies in the four L. crispatus genomes, but I did find genes similar to the glgX gene: a gene annotated as a type 1 pullulanase (uniprot link of the copy in strain MV-3A-US). One of the strains unable to grow on glycogen (125-2-CHN) had no copy of this gene and the other strain unable to grow on glycogen (JV-V01) had a mutation in the upstream region, that I thought might be a dealbreaker for expression. This paper on comparative genomics of Lactobacillus crispatus confirms the absence/presence of the type 1 pullulanase in these strains. (supplementary material file nr 5). All other strains in the comparison contained the gene except for 125-2-CHN and 214-1.
This was all purely speculative at that moment, but this is why the type 1 pullulanase gene caught my attention: it was the only one that showed clear variation amongst the L. crispatus strains. There was a big chunk of serendipity that lead me to the presented finding.
All L. crispatus strains have the type 1 pullulanase gene, but the N-terminus looks different!
I was therefore disappointed that initially, from Alicia’s dbCAN analysis, and also in a screen performed by Jorne Swanenburg, it became clear that all strains had a copy of this gene. I further analysed this gene and also included the upstream region (expecting to find a mutation similar to the one in the JV-V01 strain). The sequence directly upstream this gene encodes a putative signal peptide (see Figure 1) and there is strong variation amongst the collection of genomes in this particular area (see Figure 2)
Please find the genes from all strains here. I used the EMBL Clustal Omega online Multiple Sequence Alignment tool to compare the genes and the results were striking. You could redo the analysis by using the file and just copy paste it into the Clustal Omega tool. The genes are conserved but not the starting region. In Figure 1, a schematic overview is shown of the organization of this gene in L. crispatus. Thanks Alicia!) In Figure 2, I am showing the variation in the N-terminal sequence from the aforementioned Clustal Omega comparison.
Figure 1: Graphical Representation of GH13_13 Pullulanase Type 1. (N to C terminal): SP: signal peptide (amino acids 1-45), CBM41: carbohydrate-binding module family 41 (amino acids 110-220), CBM48: carbohydrate-binding module family 48 (amino acids 400-505), GH13_13: glycoside hydrolase family 13 subfamily 13 (amino acids 606-900), SLAP: Surface layer associated domain (1100-1259)
 Figure 2: Comparison of the N-terminal sequence of the pullulanase gene in L. crispatus strains. Adjusted from Clustal Omega. Red: strains with a possible disrupted N-terminal sequence and signal peptide. Blue: strains with an N-terminal sequence indicating an intact signal peptide. Find full sequences here.
Figure 2: Comparison of the N-terminal sequence of the pullulanase gene in L. crispatus strains. Adjusted from Clustal Omega. Red: strains with a possible disrupted N-terminal sequence and signal peptide. Blue: strains with an N-terminal sequence indicating an intact signal peptide. Find full sequences here.
7 out of the 24 strains (RL_#-strains) have a mutation in the N-terminal locus of the pullulanase gene, more specifically, in the sequence of the signal peptide. Those strains are indicated by red in Figure 2. Further experiments should indicate whether this means that the start site of these “red” strains lies more downstream from the start site in the “blue” strains.
Remarkably, there are five different variants of this gene locus present in those seven strains. For instance, strains RL_002 and RL_009 only show a deletion of two nucleotides (a frame shift), whereas strain RL_006 and RL_007 have a completely different sequence in this region.
Growth on glycogen of the L. crispatus strains.
I performed a very straightforward cultivation experiment using glycogen as a carbon source. Initially, I only used four isolates (RL_002, RL_003, RL_007 and RL_026). When these results were promising I started to screen all 23 strains . As a benchmark I used the DSM strain, which I previously showed is able to breakdown glycogen for growth and lactic acid production. I used the same methods as described in that blog post:
I inoculated the strains in regular NYCIII glucose and after ~72 hours of growth diluted them with NYCIII glycogen (final concentration 0,5%), water (as a negative control) and glucose dissolved in water (final concentration 0,5% as a positive control). After 48 hours I measured the optical density at 600 nm to determine growth on glycogen compared to growth on NYCIII without supplemented energy source and NYCIII glucose.
For 19 out of the 23 strains tested (we could not revive strain RL_005) I have either biological triplicates or duplicates, I am showing their results in Figure 3. I am still working on getting all data complete and aim to have at least two replicates of this experiment for each strain. In one biological replicate of strain RL_019 and one replicate of RL_006 results are very different from the others. I have no idea why and no ‘reason’ to exclude it. Not really sure what to do with this measurement at this point. All individual biological replicates are shown in the figure.
Note: I measured the cell density 10x diluted in PBS as well as undiluted. I am showing the diluted data in this blog post, since I do not have the undiluted culture measured on all dates for all strains. The difference between “growth” and “no growth” is more pronounced in the undiluted measurement since background absorption of the media is lower. I include an overview of the undiluted data in a separate tab of the excel sheet and the figure in the GraphPad Prism file. Other data that are in the file but not in the Figure shown here: the optical density in the positive (glucose) and negative control (water). Find the data here (.xls and the GraphPad Prism 7 file to generate the figure).
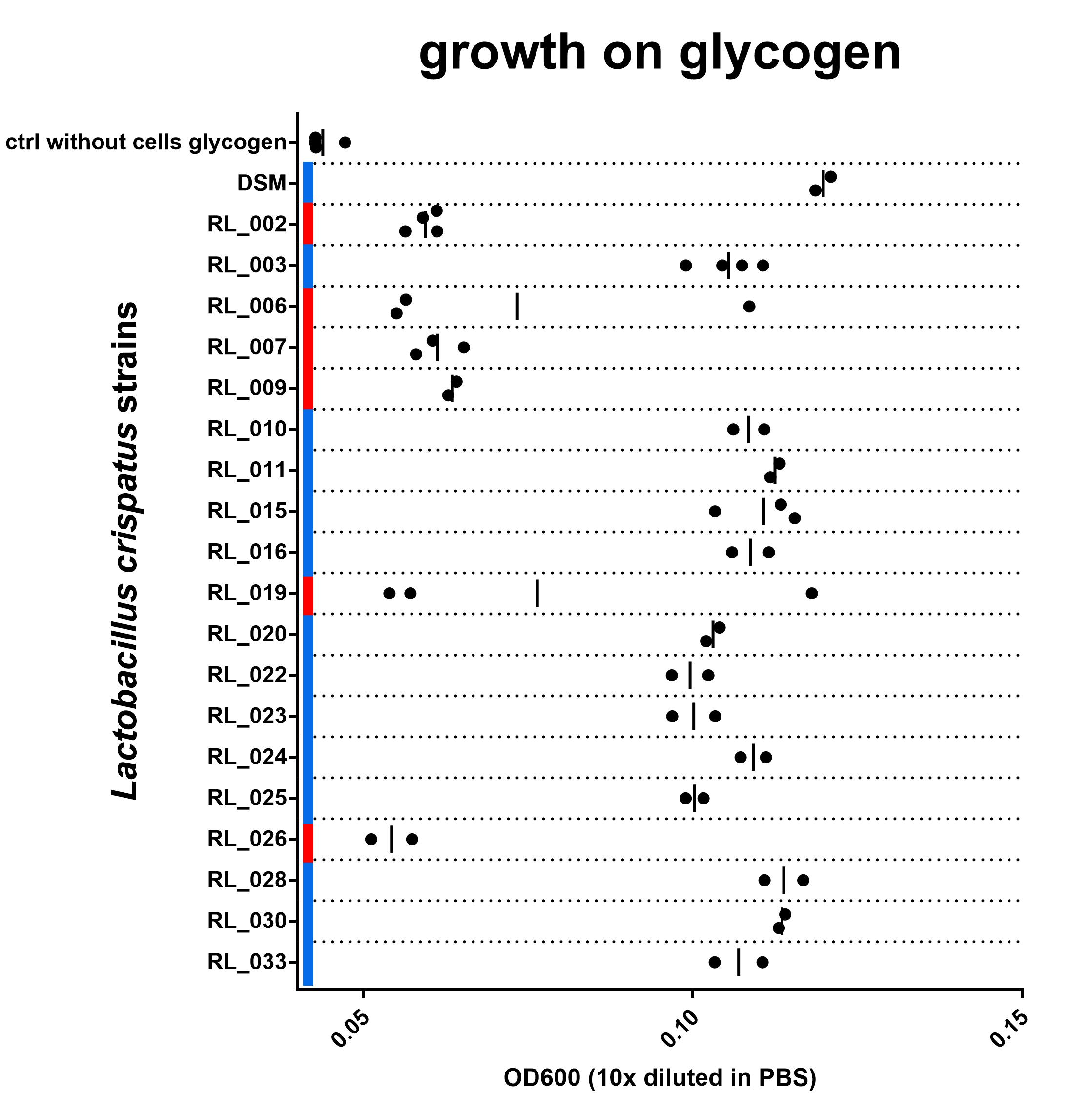
Figure 3: OD600 after culturing on NYCIIImedia with 0,5% glycogen. Biological replicates are shown as individual data points (some duplicates, some triplicates), vertical line indicates mean. Red or blue corresponds with red and blue in Figure 2. blue = with intact N-terminal signal peptide, red = with a disruption in this sequence. Find data here and protocol here
Although the growth data set is not complete yet, I do think we are seeing a strong connection between the N-terminal signal peptide of the type 1 pullulanase and the ability of the L. crispatus strain to grow on glycogen. All 6 strains in this experiment that have a disruption in the N-terminal signal peptide of the pullulanase gene show no growth on glycogen. The 14 strains that do have an intact signal peptide in the pullulanase gene can use glycogen as a source for growth. I see these data as strong evidence for an essential role of the pullulanase gene for glycogen consumption in Lactobacillus crispatus and, more specifically, the N-terminal signal peptide.
Thoughts, questions and new experimental plans
This finding is just that, a finding. Nothing final yet, an experiment that leads to a hypothesis: the N-terminal 29 amino acids are somehow important for Lactobacillus crispatus glycogen consumption. But how? Does this signal peptide lead to secretion of this enzyme? And if so, what is the influence of the C-terminal SLAP-domain? Are both required to localize this enzyme on the outer cell wall to be able to break down the big molecules of glycogen in its surroundings? Or does this signal peptide have a different function?
It is possible that the pullulanase without the signal peptide still functions in an intracellular metabolic pathway for glycogen breakdown, whereas the pullulanase gene that is localized on the outer cell wall can also debranch external glycans and utilize them for growth and lactic acid production.
I envision a few experiments to test these questions:
- it is probably important at this point to establish that the pullulanase is indeed a pullulanase. I have talked with Alicia about expressing the gene in E. coli to further characterize its activity. Alternatively, we could try to capture its native activity using cell free extracts and analyse carbohydrate products with Thin Layer Chromatography. Previously, we were able to show starch breakdown, next we should take a better look at the actual breakdown products.
- How can we study the role of the signal peptide? This is not that straightforward. Optimally we would just make L. crispatus mutants with and without it and track the enzyme’s activity and location. However, I have not seen any Lactobacillus crispatus cloning anywhere and not looking forward to try to develop my own protocols to get these isolates genetically accessible. (if someone has an idea, let me know). I hope to come up this summer with some experimental plan to test the cellular localization of this enzyme with and without the signal peptide.
What is the role of Lactobacillus crispatus glycogen metabolism in the context of the vaginal environment?
I believe that these experiments show that this activity is not something essential: the non-glycogen consumers seem to live happily in the vagina. Other lactobacilli, such as jensenii and gasseri also flourish in the vagina without being able to break down glycogen. Either, these bacteria utilize an alternative energy and carbon source. Or -what I personally expect- is that these bacteria live alongside glycogen consumers such as Lactobacillus iners or crispatus strains. Currently we don’t know whether the lactobacilli we encounter so abundantly in the vagina of reproductive age women are a collection of various strains or are a clonal population. It could well be that the non-glycogen consumers only thrive as freeloaders and can only colonize alongside a second glycogen-consuming species that does some of the glycogen breakdown. These exciting questions are definitely on my experimental wish list.
As I am still at home with baby, I will mostly do some reading and computer work to understand the genetic context and role of this pullulanase in other species. I found some interesting literature on this enzyme in Lactobacillus acidophilus and in a thermophilic bacterium called “Caldicellulosiruptor kronotskyensisencodes” (WOW! That must be one really interesting species!) Generally, pullulanases (and definitely secreted pullulanases) are of industrial relevance so there should be quite some protocols and knowledge out there. Also, I really need to read up on S-layers in lactic acid bacteria. Stay tuned!
And again: if you have suggestions how to continue, remarks or criticism, please let me know below. In general, I really appreciate any signs that this work matters to anyone because of the alternative publication route I am taking here.
Speculation on the role of bacterial peroxide production in the vaginal environment.
A new commentary on the antimicrobial role of H2O2 produced by lactobacilli in the vagina was published last week.
https://microbiomejournal.biomedcentral.com/articles/10.1186/s40168-018-0418-3
I agree with the authors, that it is not likely that H2O2 has a significant effect in keeping BV-associated bacteria at bay. Especially the experiments that dr. O’Hanlon published previously comparing the bactericidal effects of lactic acid and H2O2. They found that H2O2 is as harmful to lactobacilli as it is to several BV-associated bacteria, whereas lactobacilli can survive very high lactic acid concentrations.
I would like to add one piece of speculation here. Although the authors say that semen and cervicovaginal fluid have strong antioxidative properties that would eliminate any H2O2, I do wonder whether H2O2 production may serve a role in the onset of BV. During disruption of the vaginal environment, either through sexual arousal, intercourse, tampon insertion etc, oxygen levels are expected to rise to atmospheric levels. Is it possible that hydrogen peroxide production and accumulation in that case can result in the demise of both lactobacilli as well as BV-bacteria? This might then open up a “window of opportunity” in which general bacterial levels are temporarily low, lactic acid levels are reduced and pH is neutral. This relatively benign environment may give BV-associated bacteria a chance to colonize and proliferate.
Just a thought about a possible alternative role of H2O2.
Podcast on The Vital Question
I have been working on two things in the past months. Soon, I will post an update here on a possibly interesting gene variant I found in L. crispatus, that might be involved in glycogen metabolism.
In the meanwhile, I am proud to present to you my first podcast! The topic is “The Vital Question”, a book by Nick Lane that provides answers to the most profound question in biology “why does life look the way it does”. Game changing book according to the Guardian, and someone who agrees with that wholeheartedly is prof. Bas Teusink of the Free University of Amsterdam (also on Twitter). I sit down with Bas to talk about the beginning of the book where the author lays down “the problem” . That problem is that although life looks very diverse from the human eye, when you get down to the chemistry and the energetic systems we are all very similar.
I sincerely hope you enjoy it! Please let me know what you think either below or on Twitter. I am not exactly sure where I am headed with this, whether it’s a one time thing, or a pilot to something bigger. Your feedback is much appreciated.